Samuel Gratzl
Toolsmith for explorers of the information landscape on their treasure hunt for valuable insights
Samuel Gratzl is a passionate Research Software Engineer with a focus on interactive data exploration. He is a full-stack developer with 10+ years of experience. In 2017, he finished his PhD in Computer Science with a focus on Information Visualization at the Johannes Kepler University, Linz, Austria. He loves to dig into code, hunt bugs, and design new platforms. His goal is to enable researchers to discover more insights easier and faster, as well as develop libraries that help other developers do the same.
Skills
D3, Vega, Plot.ly, ggplot2, matplotlib, Tableau, PowerBI
Jupyter, RMarkdown, Quarto, Dash, RShiny, tidyverse, pandas, numpy
PostgresSQL, SQLServer, MongoDB, Redis, ElasticSearch, Neo4j, SQL
JavaScript/Typescript, React, Svelte, Vue
R, Python, FastAPI, REST, Swagger, OpenAPI, GraphQL
GitHub, Git, Kubernetes, GitHub Action, Google Cloud, AWS
Experience
- Being part of the research team analyzing healthcare data and providing feedback on current product developments.
- Curated datasets based on the internal Truveta Platform.
- Developed processes and templates for effective healthcare studies, speeding up our study creation
- Created dashboards for Vaccine Effectiveness and Adverse Events of COVID-19 Vaccines
- Identified data quality issues and supported clinical informatics in tracking them down.
- Initiated and specified product features for advanced researcher experience.
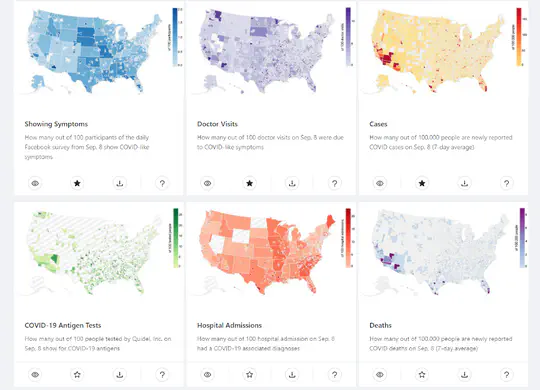
- Main front end developer of COVIDcast, a project by the Delphi Group collecting, publishing, and visualizing COVID-19 data..
- Converted the front end from a research prototype to a production-ready product.
- Enforced code quality and best practices throughout the project.
- Improved usability, maintainability, and performance of COVIDcast.
- Designed and implemented new views such as the National Survey Results View, the most popular COVIDcast view.
- Designed and implemented a new version of the COVIDcast API with increased maintainability, scalability, and robustness.
- Developed and deployed a new deployment infrastructure for the Delphi group.
- I specialize in the design and implementation of customized visual exploration web applications.
- In close collaboration with the customer, I develop specialized visual exploration platforms that not only allow the customer to answer their questions but even those they haven’t thought about yet.}
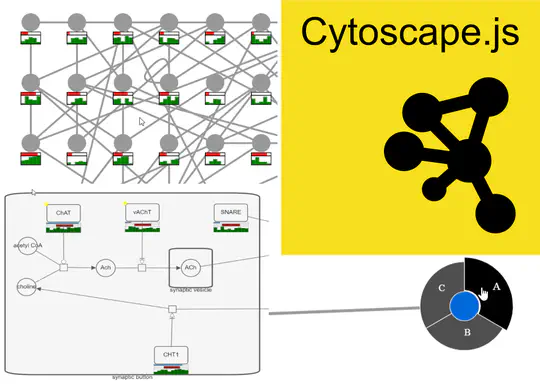
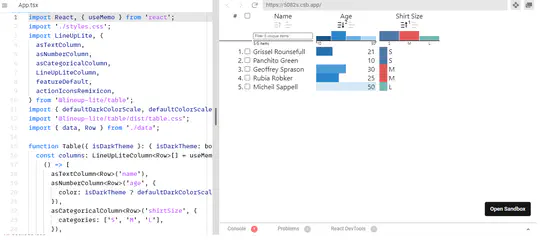
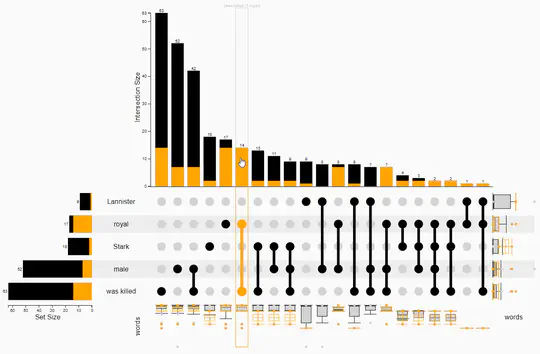
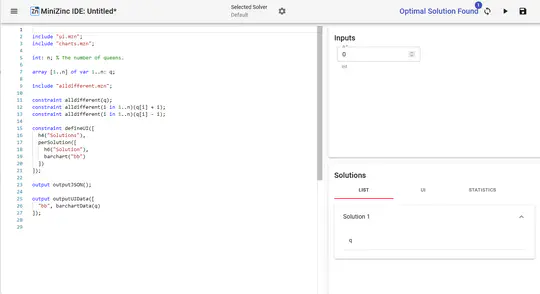
- In addition, I provide freelance service for integrating my open-source libraries, such as LinUp-lite, LineUp.js, or UpSet.js
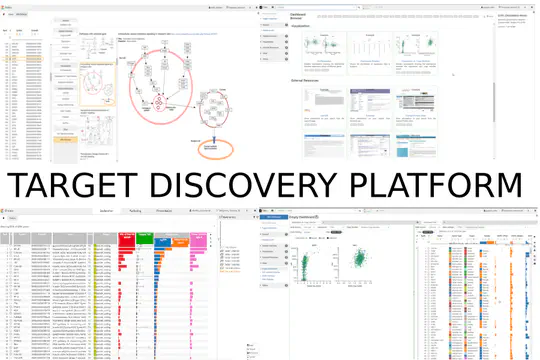
- Designed the architecture and implemented the Target Discovery Platform (TDP) with a focus on high extensibility and customizability. TDP is the foundation of all products of datavisyn and one of the three pillars of its business model.
- Built and deployed overall CI/CD infrastructure both in-house and on-premise focusing on high-availability, fault tolerance, and low maintenance.
- Lead on-site customer workshops focusing on requirements engineering, customer training, and initial prototype implementation.
- Improved project requirements to the satisfaction of the customer.
- Was the product owner for two agile customer projects which ended both in time and budget with highest customer satisfaction.
- Implemented critical features in all (4+) customer projects of datavisyn.
- Made customers happy through continuous customer support via Slack and quick response times.
- Lead, trained, and mentored the three junior developers.
- Did code reviews, introduced style guidelines, and introduced continuous testing to improve overall code quality.
- Ensured the head start of the company over competitors through integrating new technologies and frameworks.
Accomplishments
Projects
Featured Publications
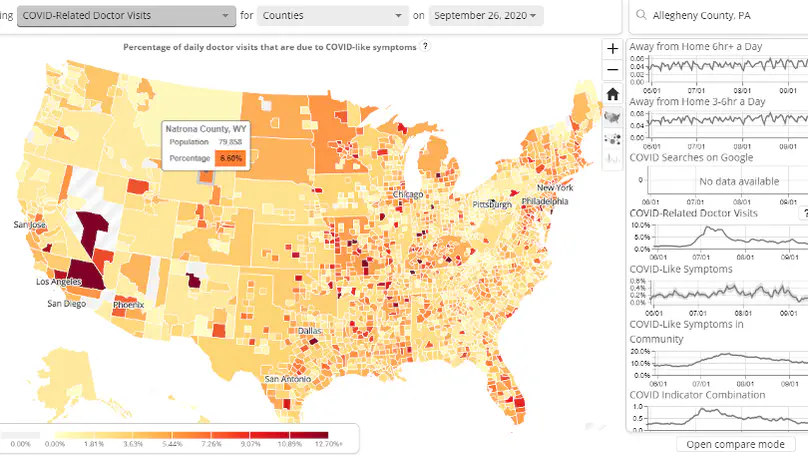
The COVID-19 pandemic presented enormous data challenges in the United States. Policy makers, epidemiological modelers, and health researchers all require up-to-date data on the pandemic and relevant public behavior, ideally at fine spatial and temporal resolution. The COVIDcast API is our attempt to fill this need: Operational since April 2020, it provides open access to both traditional public health surveillance signals (cases, deaths, and hospitalizations) and many auxiliary indicators of COVID-19 activity, such as signals extracted from deidentified medical claims data, massive online surveys, cell phone mobility data, and internet search trends. These are available at a fine geographic resolution (mostly at the county level) and are updated daily. The COVIDcast API also tracks all revisions to historical data, allowing modelers to account for the frequent revisions and backfill that are common for many public health data sources. All of the data are available in a common format through the API and accompanying R and Python software packages. This paper describes the data sources and signals, and provides examples demonstrating that the auxiliary signals in the COVIDcast API present information relevant to tracking COVID activity, augmenting traditional public health reporting and empowering research and decision-making.
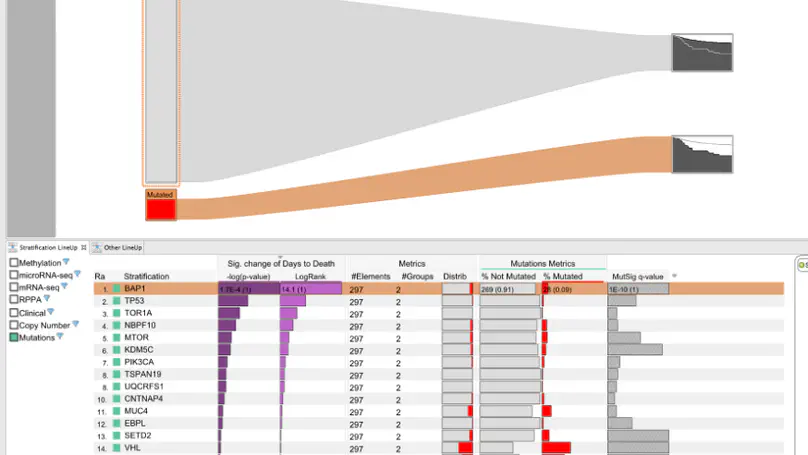
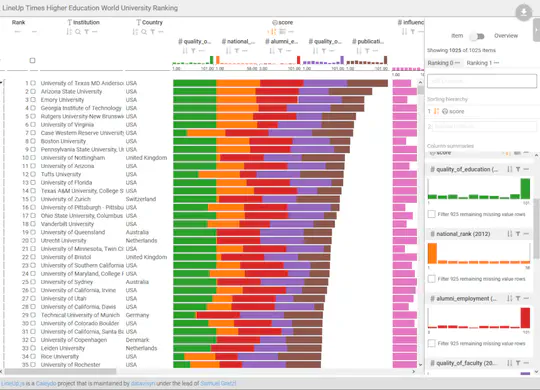
Making scientific discoveries based on large and heterogeneous datasets is challenging. The continuous improvement of data acquisition technologies makes it possible to collect more and more data. However, not only the amount of data is growing at a fast pace, but also its complexity. Visually analyzing such large, interconnected data collections requires a user to perform a combination of selection, exploration, and presentation tasks. In each of these tasks a user needs guidance in terms of (1) what data subsets are to be investigated from the data collection, (2) how to effectively and efficiently explore selected data subsets, and (3) how to effectively reproduce findings and tell the story of their discovery. On the basis of a unified model called the SPARE model, this thesis makes contributions to all three guidance tasks a user encounters during a visual analysis session: The LineUp multi-attribute ranking technique was developed to order and prioritize item collections. It is an essential building block of the proposed guidance process that has the goal of better supporting users in data selection tasks by scoring and ranking data subsets based on user-defined queries. Domino is a generic visualization technique for relating and exploring data subsets, supporting users in the exploration of interconnected data collections. Phoeva is a novel open-source visual analytics platform designed to speed up the creation of domain-specific exploration tools. The final building block of this thesis is CLUE, a universally applicable framework for capturing, labeling, understanding, and explaining visually driven exploration. Based on provenance data captured during the exploration process, users can author “Vistories”, visual stories based on the history of the exploration. The practical applicability of the guidance model and visualization techniques developed is demonstrated by means of usage scenarios and use cases based on real-world data from the biomedical domain.
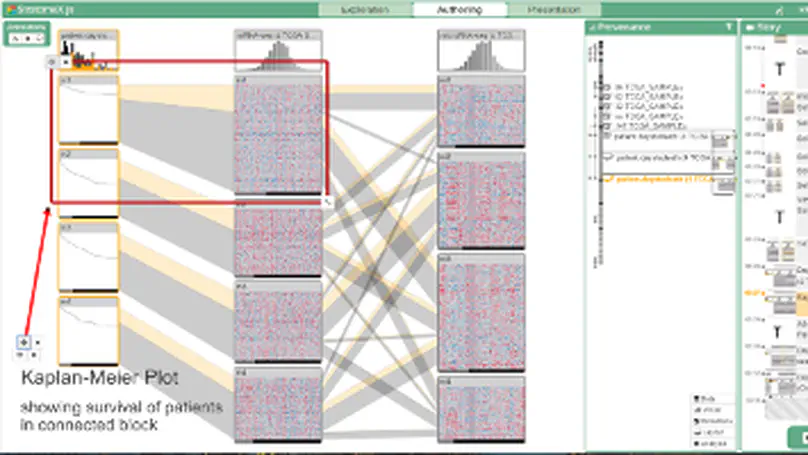
The primary goal of visual data exploration tools is to enable the discovery of new insights. To justify and reproduce insights, the discovery process needs to be documented and communicated. A common approach to documenting and presenting findings is to capture visualizations as images or videos. Images, however, are insufficient for telling the story of a visual discovery, as they lack full provenance information and context. Videos are difficult to produce and edit, particularly due to the non-linear nature of the exploratory process. Most importantly, however, neither approach provides the opportunity to return to any point in the exploration in order to review the state of the visualization in detail or to conduct additional analyses. In this paper we present CLUE (Capture, Label, Understand, Explain), a model that tightly integrates data exploration and presentation of discoveries. Based on provenance data captured during the exploration process, users can extract key steps, add annotations, and author ‘Vistories’, visual stories based on the history of the exploration. These Vistories can be shared for others to view, but also to retrace and extend the original analysis. We discuss how the CLUE approach can be integrated into visualization tools and provide a prototype implementation. Finally, we demonstrate the general applicability of the model in two usage scenarios: a Gapminder-inspired visualization to explore public health data and an example from molecular biology that illustrates how Vistories could be used in scientific journals.
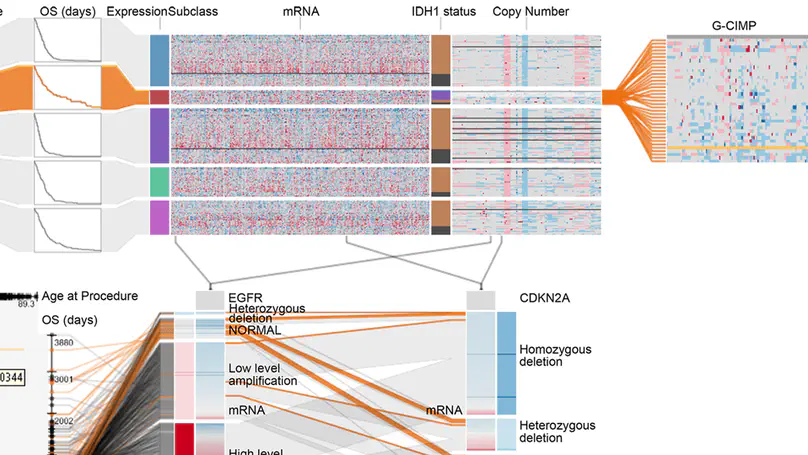
Answering questions about complex issues often requires analysts to take into account information contained in multiple interconnected datasets. A common strategy in analyzing and visualizing large and heterogeneous data is dividing it into meaningful subsets. Interesting subsets can then be selected and the associated data and the relationships between the subsets visualized. However, neither the extraction and manipulation nor the comparison of subsets is well supported by state-of-the-art techniques.
In this paper we present Domino, a novel multiform visualization technique for effectively representing subsets and the relationships between them. By providing comprehensive tools to arrange, combine, and extract subsets, Domino allows users to create both common visualization techniques and advanced visualizations tailored to specific use cases. In addition to the novel technique, we present an implementation that enables analysts to manage the wide range of options that our approach offers. Innovative interactive features such as placeholders and live previews support rapid creation of complex analysis setups. We introduce the technique and the implementation using a simple example and demonstrate scalability and effectiveness in a use case from the field of cancer genomics.
Recent Publications
Contact
- contact@sgratzl.com
- Marlton, NJ 08053